The CMCF Mathematics Core focuses on method development to address gaps in our ability to predict, infer, and model multiscale biological systems. These are closely interlinked, and in addressing the gaps we emphasize new tools that enable analysis of individual cells by connecting genes and cell states through ‘lenses’ of varying magnification. Another focus is on spatiotemporal integration at gene-cell-tissue scales for new multiscale models of cell fate.
We are developing tools for the analysis of single-cell RNA sequencing (scRNA-seq) data to address challenges such as the lack of consistency between tasks (e.g. clustering vs. pseudotime inference), deviations due to noise, variability, and batch effects, gene-gene or cell-cell dependencies, and methods for spatial inference.
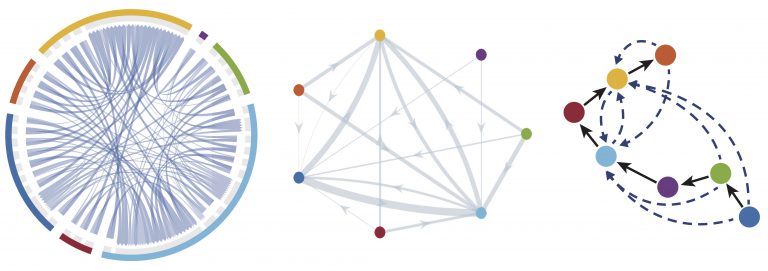
We develop stochastic algorithms for gene regulatory network modeling to better study the variability within and between cells. By modeling gene regulatory networks at the single-cell level, we also take into account differences between cell types, heterogeneity in gene expression within a single cell population, cell division, and transitions between different states. Major challenges of these approaches include stiffness due to multiple temporal scales, characterizing rare but important cellular states, identifying irreversible or discrete state transitions, and the incorporation of epigenetic factors. As we move up scales to multicellular systems, single cell resolution is required to dissect heterogeneous cell fates both in terms of function and genotype, thus requiring approaches such as the Subcellular Element Method (SEM) to model these systems. SEM describes a number of elements connected by subcellular forces, thus capturing details such as cell polarity, cell-cell contact, and basic cell shape, while remaining computationally tractable for the simulation (in 2D and in 3D) of large numbers of cells.
Often, model parameters are impossible or infeasible to measure directly. To address this we develop Bayesian frameworks to link models with measurements, accounting both for prior knowledge of the systems and for our uncertainties. We focus on developing statistical inference approaches for modeling with hierarchical structure and non-linear relationships among variables, to understand the relationships between biological processes across multiple scales. Concurrently, we are developing methods for parameter estimation using hybrid variational and Hamiltonian Monte Carlo methods, to provide sufficient computational power for inference.
